Numpy
NumPy is a successor to the Numeric array object, aiming to establish a foundation for scientific computing. The author, a graduate student in biomedical imaging, discovered Python and Numeric in 1998 and became involved in the Numeric Python community. Pearu Peterson’s development of f2py facilitated wrapping Fortran programs into Python, contributing to scientific computing with Python. In 2001, SciPy was formed through the collaboration of the author, Eric Jones, and Pearu Peterson, combining Python modules for scientific computing. Numarray was created as a replacement for Numeric, leading to some fragmentation in the Python scientific computing community. The author initiated efforts to bring the community together, ultimately resulting in the development of NumPy, released in 2006. NumPy 1.0 was released in late 2006, offering enhanced universal functions and features, and the package adopted the NumPy name.
| Sub-package | Purpose | Comments |
|---|---|---|
| core | basic objects | all names exported to numpy |
| lib | additional utilities | all names exported to numpy |
| linalg | basic linear algebra | old LinearAlgebra from Numeric |
| fft | discrete Fourier transforms | old FFT from Numeric |
| random | random number generators | old RandomArray from Numeric |
| distuils | enhanced build and distribution | improvements built on standard disutils |
| testing | unit-testing | utility functions useful for testing |
| f2py | automatic wrapping of Fortan code | a useful utility needed by SciPy |
NumPy provides two fundamental objects: an N-dimensional array object (ndarray) and a universal function object (ufunc). An N-dimensional array is a homogeneous collection of items indexed using N integers. There are two essential pieces of information that define an N-dimensional array:
- the shape of the array
- the kind of item the array is composed of
np.array(object=, dtype=None, copy=True)
An ndarray is an N-dimensional array of items where each item takes up a fixed number of bytes. Typically, this fixed number of bytes represents a number (e.g. integer or floating-point). However, this fixed number of bytes could also represent an arbitrary record made up of any collection of other data types. NumPy achieves this flexibility through the use of a data-type (dtype) object. Every array has an associated dtype object which describes the layout of the array data. Every dtype object, in turn, has an associated Python type-object that determines exactly what type of Python object is returned when an element of the array is accessed.
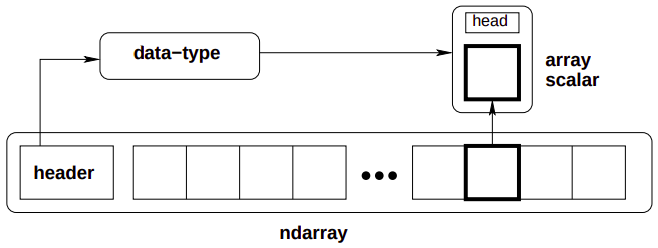
Memory layout:
Each of these arrays take 12 blocks of memory. How this memory is used to form the abstract 2-dimensional array can vary, however, the ndarray object supports both style.
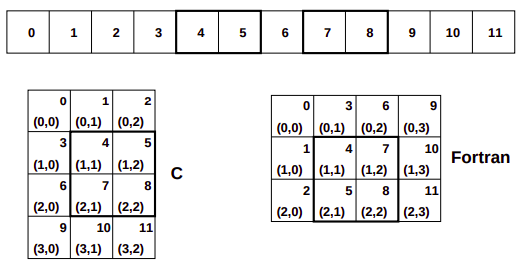
In the C-style of N-dimensional indexing, the last N-dimensional index varies the fastest. In other words, to move through computer memory sequentially, the last index is incremented first, followed by the second-to-last index and so forth. In the Fortran-style of N-dimensional indexing, the first N-dimensional index varies the fastest. Thus, to move through computer memory sequentially, the first index is incremented first until it reaches the limit in that dimension, then the second index is incremented and the first index is reset to zero. The styles of memory layout for arrays are connected through the transpose operation. Thus, if is a C-style array, then the same block of memory can be used to represent as a (contiguous) Fortran-style array.
(arrays are stored as what (C or Fortan) contiguous)
Let be the value of the th index into an array whose shape is represented by the integers . The formulas for the one-dimensional index of the N-dimensional arrays reveal what results in an important generalization for memory layout. Note that each formula can be written as: where gives the stride for dimension . Thus, for and Fortan contiguous arrays respectively we have: As long as we always use the stride information to move around in the N-dimensional array, we can use any convenient layout we wish for the underlying representation as long as it is regular enough to be defined by constant jumps in each dimension. The ndarray object of NumPy uses this stride information and therefore the underlying memory of an ndarray can be laid out dis-contiguously. An important situation where irregularly strided arrays occur is array indexing.
Universal functions:
The ufunc is an instance of a general class so that function behavior is same. All ufuncs perform element by element operations over an array or a set of arrays (for multi-input functions). An important aspect of ufunc is the idea of broadcasting. Broadcasting allows ufuncs to deal in a meaningful way with inputs that do not have exactly the same shape. In particular,
- The first rule of broadcasting is that if all input arrays do not have the same number of dimensions, then a 1 will be prepended to the shapes of the smaller arrays until all the arrays have the same number of dimensions.
- The second rule of broadcasting ensures that arrays with a size of 1 along a particular dimension act as if they had the same size of the array with the largest shape along that dimension. The value of the array element is assumed to be the same along that dimension for the broadcasted array.
The most common alternation needed is to route-around the automatic prepending of 1’s to the shape of the array. If it is desired, to add 1’s to the end of the shape, then dimension can be added using np.newaxis name in NumPy.
Application:
One important aspect of broadcasting is the calculation of functions on regularly spaced grids. For example, suppose it is desired to show a portion of the multiplication table by computing the function a*b on a grid with a and b running from.
a[:,newaxis]*bThe Array Attributes:
Array attributes reflect information that is intrinsic to the array itself. The exposed attributes are the core parts of an array and only some of them can be reset meaningfully without creating a new array.
| Attribute | Description |
|---|---|
| flags | dictionary-like attributes showing the state of flags in this array |
| ndim, shape | number of dimensions in array, array shape tuple |
| size, itemsize, nbytes | number of elements, element size, number of bytes |
| dtype | data-type object for this array |
| strides | bytes to jump in the data segment to get from one entry to next |
In NumPy, the ndarray object has many methods which operate on or with the array in some fashion, typically returning an array result.
The Array Methods:
The ndarray object has many methods which operate on or with the array in some fashion, typically returning an array result.
Array conversion methods:
| Method | Arguments | Description |
|---|---|---|
| astype | (dtype {None}) | Cast to another data type |
| copy | () | Copy array |
| tolist | () | Array as a nested list |
| dump | (file) | Pickle to stream or file |
| dumps | () | Gets pickled string |
Array shape manipulation:
| Method | Arguments | Description |
|---|---|---|
| reshape | (newshape, order=‘C’) | Return an array that uses same data but new shape |
| transpose | () | Return an array view with the shape transposed |
| flatten | () | Return a new 1-d array with data copied from self |
| ravel | () | Return a 1-d version of self |
Array item selection and manipulation:
Basic indexing:
Indexing is a powerful tool in Python and NumPy takes full advantage of this power. There are three differences:
- slicing can be done over multiple dimensions
- exactly one ellipsis object can be used to indicate several dimensions at once
- slicing cannot be used to expand the size of an array (unlike lists)
There are two kinds of indexing available using X[obj] syntax: basic slicing, and advanced indexing, where X is the array to-be-sliced and obj is the selection object. These two methods of slicing have different behavior and are triggered depending on obj.
Basic slicing occurs when obj is a slice object (constructed by start:stop:step) notation inside of brackets), an integer, or a tuple of slice objects and integers. Ellipsis and newaxis object interspersed with these as well. In Python X[(exp1, exp2, ..., expN)] is equivalent to X[exp1, exp2, ...., expN] as the latter is just syntactic sugar for the former.
- The basic slice syntax is
i:j:kwhere is the starting index, is the stopping index, and is the step . - An integer, , returns same values as
i:i+1except the dimensionality of the returned object is reduced by1. - You may use slicing to set values in the array, but unlike lists you can never grow the array. The size of the value to be set in
X[obj] = valuemust be broadcastable to the same shape asX[obj].
Advanced selection: ((WHAT THE?))
Advanced selection is triggered when the selection object, obj is
- a non-tuple sequence object,
- an ndarray (of data type integer or bool), or
- a tuple with at least one sequence object or ndarray (of data type integer or bool). There are two types of advanced indexing: integer and Boolean. Advanced selection always returns a copy of the data (contrast with basic slicing that returns a view.)
…
The advanced selection occurs when obj is an array object of Boolean type (such as may be returned from comparison operators). The special case when obj.ndim == X.ndim is worth mentioning. In this case, X[obj returns a 1-dimensional array filled with th elements of X corresponding to the True values of obj. The search order will be C-style (last index varies the fastest).
Array object calculation methods:
Many of the methods take an argument named axis. In such cases, if axis is None (the default), the array is treated as a 1-d array and the operation is performed over the entire array. If axis is an integer, then the operation is done over the given axis (for each 1-d subarray that can be created along the given axis). The parameter dtype specifies the data type over which a reduction operation (like summing) should take place.
| Method | Arguments | Description |
|---|---|---|
| max, min, mean | (axis=None) | maximum/minimum/mean of self |
| sum, prod | (axis=None) | add/multiply elements of self together |
| var, std | (axis=None) | variance/standard deviation of self |
| all, any | (axis=None) | true if all/any entries are true |
| argmax, argmin | (axis=None) | index of largest/smallest value |
| clip | (min=, max=) | self[self>max]=max; self[self<min]=min |
Basic routines:
Creating arrays:
| Routine | Arguments |
|---|---|
| arange | (start=, stop=None, step=1, dtype=None) |
| linspace | (start=, stop=, num=50) |
| zeros | (shape=, dtype=int) |
| zeros_like | (arr) |
| ones | (shape=, dtype=int) |
| ones_like | (arr) |
| identify | (n, dtype=intp) |
| where | (condition[, x, y]) |
where: Returns an array shaped like condition, that has elements of x and y respectively where condition is respectively true or false. If x and y are not given, then returns n-dimensional index for elements of the n-dimensional array self that meet the condition into an n-tuple of equal-length index arrays.
(where prepares for advanced indexing i guess)
Operations on two or more arrays:
| Routine | Arguments | Remarks |
|---|---|---|
| inner | (a, b) | |
| dot | (a, b) | |
| matmul | (a,b) | (tensor contraction) |
| outer | (a, b) | A.ravel()[:,newaxis] x B.ravel()[newaxis,:] |
| convolve | (x, y, mode=‘valid’) | polynomial multiplication for 1-d arrays |
| concatenate | (seq=, axis=0) | must have compatible types |
| vstack | (seq) | stack a sequence of arrays along the first axis |
| hstack | (seq) | stack a sequence of arrays along the second axis |
| einsum | (subscripts, *operands) |
inner vs dot:
The inner product between two arrays is an array that has shape a.shape[:-1]+b.shape[:-1] with elements computed as the sum of the product of the elements from the last dimensions of a and b. In particular, let I and J be the super indices selecting the 1-dimensional arrays a[I,:] and b[J,:], the the resulting array, r, is:
Ordinary inner product of vectors for 1-D arrays, in higher dimensions a sum product over the last axes.
The dot product between two arrays is product-sum over the last dimension of a and the second-to-last dimension of b. Specially, if I and J are super indices for a[I,:] and b[J,:,j] so that j is the index of the last dimension of b. Then shape of the resulting array is a.shape[:-1]+b.shape[:-2]+b.shape[-1] with elements:
For 2-D arrays it is equivalent to matrix multiplication, and for 1-D arrays to inner product of vectors. For N dimensions it is a sum product over the last axis of a and the second-to-last axis of b.
hstack vs vstack:
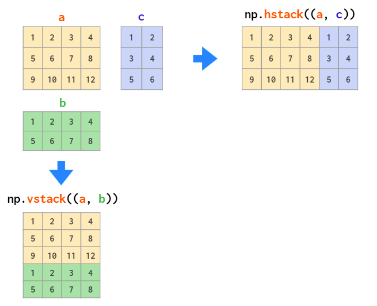
The OG einsum:
Suppose we have two arrays, A and B. Now we want to :
- multiply
AandBin a particular way to create a new array of products, and then maybe - sum this new array along particular axes, and/or
- transpose the axes of the array in a particular order Then there is a good chance einsum will help us do this much faster and more memory-efficiently that combinations of the NumPy functions multiply, sum and transpose would allow.
The key is to choose the correct labelling for the axes of the inputs arrays and the array that we want to get out. A good example to look at is matrix multiplication, which involves multiplying rows with columns and then summing the products. For two 2D arrays A and B, matrix multiplication can be done with:
np.einsum('ij,jk->ik', A, B)The left-hand part of the labels the axes of the input arrays: ij labels A and jk labels B. The right-hand part of the string labels the axes of the single output array with the letters ik. Drawing on the labels:
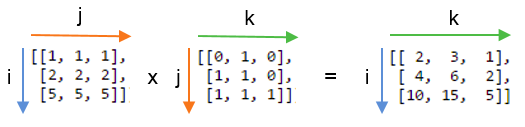
Repeating letters between input arrays means that values along those axes will be multiplied together. The products make up the values for the output array.
Omitting a letter from the output means that values along that axis will be summed.
We can return the unsummed axes in any order we like.
If we didn’t sum the j axis and instead included it in the output by writing `np.einsum(“ij,jk→ijk”, A,B):
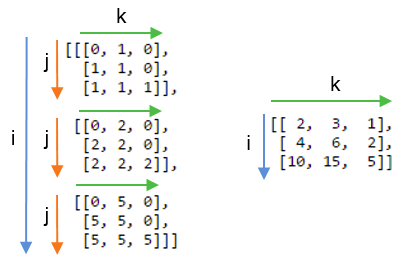
Basic calculus functions:
| Routines | Arguments | Remarks |
|---|---|---|
| histogram | (x=, bins=None) | |
| diff | (x, n=1, axis=-1) | Calculates nth order difference along given axis |
| gradient | (f, varags, axis=None) | Returns tuple (n, bins) where n is the histogram |
| trapz | (y, x=None, dx=1.0, axis=-1) |
The gradient routine uses central differences on the interior and first differences on boundaries to give the same shape for each component of the gradient. The varags variable can contain 0, 1, or N scalar corresponding to the sample distances in each direction (default 1.0). If f is N-d, then N arrays are returned each of the same shape as f, giving the derivative of f with respect to each dimension.
For derivative:
x = np.linspace(0, 10, 1000)
y = f(x)
dx = x[1]-x[0]
dydx = np.gradient(y, dx)For a tabular value x and y, find all the local minimas with:
np.where(np.diff(np.sign(dydx))>0)[0] #indices of minimasIf y contains samples of a function: yi = f(xi) then trapz can be used to approximate integral of the function using trapezoidal rule. If the sampling is not evenly spaced use x to pass in the sample positions. Otherwise, only the sample-spacing is needed in dx. The trapz function can work with many functions at a time stored in an N-dimensional array. The axis argument controls which axis defines the sampling axis (the other dimensions are different functions). The number of dimensions of the returned result is y.ndim - 1.
For a tabular value x and y, find integral with:
dx=x[1]-x[0]
np.trapz(y, dx=dx)Universal functions:
Universal functions are wrappers that provide a common interface to mathematical functions that operate on scalars, and can be made to operate on arrays in an element-by-element by fashion. All ufuncs wrap some core function that takes in ni (scalar) inputs and produces n0 (scalar) outputs. Typically, this core function is implemented in compiled code but a Python function can also be wrapped into a universal function using the basic method frompyfunc in the umath module.
The standard broadcasting rules are applied so that inputs without exactly the same shapes can still be usefully operated on.
Internally, buffers are used for misaligned data, and data that has to be converted from one data type to another. The size of the internal buffers is settable on a per-thread basis. There can be up to 2(ni+n0) buffers of the specified size created to handle the data from all the inputs and outputs of a unfunc. The default size of the buffer is 10,000 elements. Whenever buffer-based calculation would be needed, but all input arrays are smaller than the buffer size, those misbehaved or incorrect typed arrays will be copied before the calculation proceeds.
Methods:
All ufuncs have 4 methods. However, these methods only make sense on ufuncs that take two input arguments and return one output argument. Attempting to call these methods on other ufuncs will cause a ValueError. The reduce-like methods all take an axis keyword and a dtype keyword, and the arrays must all have dimension >= 1. The axis keyword specifies which axis of the array the reduction will take place over and may be negative, but must be an integer. The dtype keyword allows you to manage a very common problem that arises when naively using .reduce.
| Method | Arguments | Description |
|---|---|---|
| reduce | (array=, axis=0, dtype=None) | |
| accumulate | (array=, axis=0, dtype=None) | returns an array of the same shape |
| outer | ||
| reduceat |
Random:
The fundamental random number generator is the Mersenne Twister based on code written by Makoto Matsumoto and Takuji NIshimura (and modified for Python by Raymond Hettinger). Random numbers from discrete and continuous distributions are available, as well as some useful random-number-related utilities. Each of the discrete and continuous random number generators take a size keyword. If this is None (default), then the size is determined from the additional inputs (using ufunc-like broadcasting). If no additional inputs are needed, or if these additional inputs are scalars, then a single number is generated from the selected distribution. If size is an integer, then a 1-d array of that size is generated filled with random numbers from the selected distribution. Finally, if size is a tuple, then an array of that shape is returned filled with random numbers.
Discrete distributions:
| Distribution | Arguments | Description |
|---|---|---|
| binomial | (n, p, size=None) | This random number models the number of successes in n independent trials of a random experiment where the probability of success in each experiment is p. |
| geometric | (p, size=None) | This random number models the number of (independent) attempts required to obtain a success where the probability of success on each attempt is p. |
| randint | (low, high=None, size=None) | Equally probable random integers in the given range low ⇐ x < high. If high is None, then th range is 0 ⇐ x < low |
| hypergeometric | (ngood, nbad, nsample, size=None) | There are two types of objects in a jar. The hypergeometric random number models how many good objects will be present when N items are taken out of the urn without replacement |
Continuous distributions:
| Distribution | Arguments | Description |
|---|---|---|
| normal | (loc=0.0, scale=1.0, size=None) | The normal distribution is limiting distribution of independent samples from any sufficiently well behaved distributions. |
| uniform | (low=0.0, high=1.0,size=None) | Returns random numbers that are equally probable over the range low, high |
| rand(n) | (d1, d2, …, dn) | A convenient interface to obtain an array of shape (d1, d2, …, dn) of uniform/normal random numbers in the interval |
Others:
| Routine | Arguments | Description |
|---|---|---|
| choice | (arr, size=None, p=None) | Generates a random sample from a given 1-D array. |
| shuffle | (arr) | Modify a sequence in-place by shuffling its contents. |
Linear algebra:
Fast fourier transform:
Pandas
Fundamentally, data alignment is intrinsic. The link between labels and data will not be broken unless done so explicitly by you. Series is a one-dimensional labeled array capable of holding any data type (integers, strings, floating point numbers, Python objects, etc.). This axis labels are collectively referred to as the index. The basic method to create a Series is to call.
s = pd.Series(data, index=index)Here, data can be many different things:
| Data | what index is |
|---|---|
| scalar | value will be repeated to match the length of index |
| dict | if an index is passed, the values in data corresponding to labels in the index will be pulled out |
| ndarray | same length as data, if no index is passed, one will be created having values |
| Series is ndarray-like: | |
| Series acts very similarly to a ndarray and is a valid argument to most NumPy functions. However, operations such as slicing will also slice the index. Like a NumPy array, a pandas Series has a single dtype. This is often a NumPy dtype. However, pandas extend NumPy’s type system in a few places. Some examples within pandas are Categorical data and Nullable integer data type. |
Series is dict-like: A Series is also like a fixed-size dict in that you can get and set values by index label.
DataFrame: DataFrame is a 2-dimensional labeled data structure with columns of potentially different types. You can think of it like a dict of Series objects. Like Series, a DataFrame accepts many different kinds of input.
…
Attributes:
| Attributes | Description |
|---|---|
| index | The index (row labels) of the DataFrame. |
| columns | The column labels of the DataFrame. |
| dtypes | Return the dtypes in the DataFrame. |
| memory_usage | Return the memory usage of each column in bytes. |
Indexing/selection:
| Operation | Syntax | Result |
|---|---|---|
| Select column | df[col] | Series |
| Select multiple columns | df[[col1, col2, ..., coln]] | DataFrame |
| Select row by label | df.loc[label] | Series |
| Select row by integer location | df.iloc[loc] | Series |
| Slice rows | df[0:5] | DataFrame |
| Select rows by boolean vector | df[bool_vec] | DataFrame |
Iteration:
| Methods | Description |
|---|---|
| Iterate over (column name, Series) pairs | df.keys() |
| Iterate over DataFrame rows as (index, Series) pairs | df.iterrows() |
Data structures:
Series:
s.index
s.values
dictt-like: ['key'] -> dtype; [['key', 'key']] -> series;
array-like: iloc['idx'] -> dtype; ['idx':'idx'] -> series;
Dataframe:
pd.DataFrame(series_of_dict)
pd.DataFrame(dicts_of_array-like)
df.index
df.values
df.columns
dictt-like: ['col_key'] -> series, [['col_key', 'col_key']] -> dataframe;
array-like: iloc['idx'] -> series, ['idx':'idx'] -> dataframe;
array-like: [df.index.get_loc('row_key')];
Methods:
dfs.head(n=5)
dfs.tail(n=5)
dfs.info()
dfs.to_numpy()Matplotlib
Implicit plot: matlab like way of plotting
Explicit axes: instantiating an instance of Figure class, using a subplots method or similar on that object to create one or more Axes objects and then calling drawing methods on the Axes.
Creation:
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(fh, fw))
fig.suptitle("Main Title")
fig.tight_layout()Customization:
ax.cla()
ax.plot([x1], y, [fmt], x[n], y, [fmt])
fmt = [marker][line][color]
ax.legend(['Label1', 'Labeln'])
values, counts = np.unique(x, return_counts=True)
ax.bar(values, counts)
ax.hist(x, bins=None)
ax.scatter(x, y, s=None, c=None)
ax.imshow(X, cmap=None, alpha=None, interpolation=None)
ax.set_title("Axes Title")
ax.set_xlabel(xlabel)
ax.set_ylabel(ylabel)Imaging Library
PyTorch
Tensors are a generalization of vectors and matrices. In PyTorch, they are a multi-dimensional matrix containing elements of a single data type. They have a type, a shape and live in some device.
Although PyTorch has an elegant python first design, all PyTorch heavy work is actually implemented in C++. In Python, the integration of C++ code is (usually) done using what is called an extension. PyTorch uses ATen, which is foundational tensor operation library on which all else is built. To do automatic differentiation, PyTorch uses Autograd, which is an augmentation on top of the ATen framework.
It is very common to load tensors in numpy and convert them to PyTorch, or vice-versa;
np_array = np.ones((2,2))
torch_array = torch.tensor(np_array)
torch_array = torch.from_numpy(np_array)Difference between in-place and standard operations might not be so clear in some cases:
#in-place:
torch_array += 1.0
torch_array.add_(1.0)
#standard:
torch_array = torch_array + 1.0 Tensor storage:
The abstraction responsible for holding the data isn’t actually the Tensor, but the Storage. It holds a pointer to the raw data and contains information such as the size and allocator. Storage is a dump abstraction, there is no metadata telling us how to interpret the data it holds.
struct C10_API StorageImpl : public c10::intrusive_ptr_target {
...
DataPtr data_ptr_;
SymInt size_bytes_;
Allocator* allocator_;
}The Storage abstraction is very powerful because it decouples the raw data and how we can interpret it. We can have multiple tensors sharing the same storage, but with different interpretations, also called views, but without duplicating memory.
x = torch.ones((2,2))
x_view = x.view(4)
x_data = x.untyped_storage().data_ptr()
x_view_data = x_view.untyped_storage().data_ptr()
x_data == x_view_data #TrueMemory allocators:
The tensor storage can be allocated either in the CPU memory or GPU, therefore a mechanism is required to switch between these different allocations. There are allocators that will use the GPU allocators such as cudaMalloc() when the storage should be used for the GPU or posix_memalign() POSIX functions for data in the CPU memory.
PyTorch uses a CUDA caching allocators that maintains a cache allocations with the Block structure. The torch.cuda.empty_cache() will release all unused blocks.
The Tensor has Storage which in turn has a pointer to the raw data and to the Allocator to allocate memory according to the destination device.
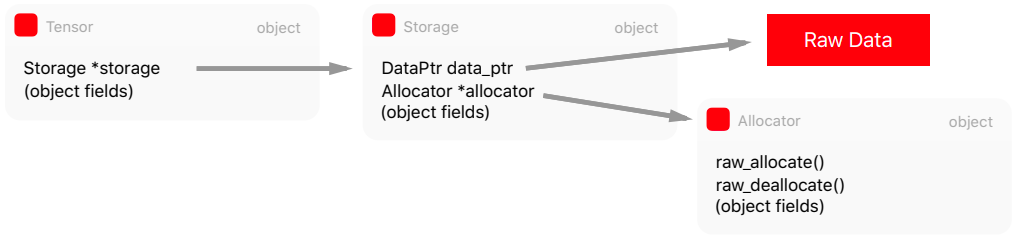
Just-in-time compiler:
PyTorch is eager by design, which means that it is easily hackable to debug, inspect etc. However, this poses problems for optimization and for decoupling it from Python (the model itself is a Python code). PyTorch introduced torch.jit, which has two main methods to convert a PyTorch model to a serializable and optimizable format.
(and others)
Tensors: ndarrays that run on GPUs
The torch.tensor() routine always copies data. If you have a Tensor and want to change its requires_grad flag, use requires_grad_() or detach() to avoid a copy. If you have a numpy array and want to avoid a copy, use torch.as_tensor(). The detach returns a new Tensor, detached from the current graph.
Autograd engine:
torch.autograd is PyTorch’s automatic differentiation engine that powers neural network training. Neural networks (NNs) are a collection of nested functions that are executed on some input data. These functions are defined by parameters (consisting of weights and biases), which in PyTorch are stored in tensors.
Training a NN happens in two steps:
Forward propagation: In forward prop, the NN makes it best guess about the correct output. It runs data through each of its functions to make this guess.
Backward propagation: In backprop, the NN adjusts its parameters proportionate to the error in its guess. It does this by traversing backwards from the output, collecting the derivatives of the error with respect to the parameters of the functions (gradients), and optimizing the parameters using gradient descent.
Example:
We create two tensors a and b with requires_grad=True which signals to autograd that every operation on them should be tracked.
a = torch.tensor([2., 3.], requires_grad=True)
b = torch.tensor([6., 4.], requires_grad=True)We create another tensor Q from a and b.
Q = 3*a**3 - b**2When we call .backward() on Q, autograd calculates gradients and stores them in respective tensors’ .grad attribute. We need to explicitly pass a gradient argument in Q.backward() because it is a vector. gradient is a tensor of the same shape as Q, and it represents the gradient of Q w.r.t itself,
Under the hood, each primitive autograd operator is really two functions that operate on Tensors. The forward function computes output Tensors from input Tensors. The backward function receives the gradient of the output Tensors with respect to some scalar value, and computes the gradient of the input Tensors with respect to that same scalar value.
In PyTorch we can easily define our own autograd operator by defining a subclass of torch.autograd.Function and implementing the forward and backward functions. We can then use our new autograd operator by constructing an instance and calling it like a function, passing Tensors containing input data.
external_grad = torch.tensor([1., 1.]) # of Q wrt to a scalar;
Q.backward(gradient=external_grad)Equivalently, we can also aggregate Q into a scalar and call backward implicitly, like:
Q.sum().backward()Vector calculus:
Mathematically, if you have a vector function essentially a NN, then gradient of with respect to is a Jacobian matrix :
J= \left(\begin{array}\\ \frac{\partial \vec{y}}{\partial x_1} & \ldots & \frac{\partial \vec{y}}{\partial x_n}\\\end{array}\right) = \left(\begin{array}\\ \frac{\partial y_1}{\partial x_1} & \ldots & \frac{\partial y_1}{\partial x_n} \\ \vdots & \ddots & \vdots\\ \frac{\partial y_m}{\partial x_1} & \ldots & \frac{\partial y_m}{\partial x_n}\end{array} \right) $$ Generally `torch.autograd` is an engine for computing vector-Jacobian product. That is, given any vector $\vec{v}$, compute the product $J^T\cdot \vec{v}$ . This characteristics of vector-Jacobian product is what we use in above example; `external_grad` represents $\vec{v}$. *Conceptually, autograd keeps a record of data (tensors) and all executed operations (along with the resulting new tensors) in a directed acyclic graph (DAG) consisting of `Function` objects. In this DAG, leaves are the input tensors, roots are the output tensors. By tracing this graph from roots to leaves, you can automatically compute the gradients using the chain rule.* In a forward pass, autograd does two things simultaneously: - run the requested operation to compute a resulting tensor, and - maintain the operation's gradient function in the DAG The backward pass kicks off when `.backward()` is called on the DAG root. `autograd` then: - compute the gradient from each `.grad_fn` - accumulates them in the respective tensor's `.grad` attribute, and - using the chain rule, propagate all the way to the leaf tensors. ![[Pasted image 20231223213121.png]] DAGs are dynamic in PyTorch: An important thing to note is that the graph is created from scratch; after each `.backward()` call, autograd starts populating a new graph. This is exactly what allows you to use control flow statements in your model; you can change the shape, size and operations at every iteration if needed. **Exclusion from the DAG:** `torch.autograd` tracks operations on all tensors which have their `requires_grad` flag set to `True`. For tensors that don't require gradients, setting this attribute to `False` excludes it from the gardient computation DAG. The output tensor of an operation will require gradients even if only a single input tensor has `requires_grad=True` In a NN, parameters that don’t compute gradients are usually called **frozen parameters**. It is useful to “freeze” part of your model if you know in advance that you won’t need the gradients of those parameters (this offers some performance benefits by reducing autograd computations). **Extending PyTorch:** Adding operations to autograd requires implementing a new Function subclass for each operation. Recall that Functions are what autograd uses to encode the operation history and compute gradients. ```Python class LinearFunction(Function): @staticmethod def forward(input, weight, bias): output = input.mm(weight.t()) if bias is not None: output += bias.unsqueeze(0).expand_as(output) return output @staticmethod def backward(ctx, grad_output): input, weight, bias = ctx.saved_tensors grad_input = grad_weight = grad_bias = None if ctx.needs_input_grad[0]: grad_input = grad_output.mm(weight) if ctx.needs_input_grad[1]: grad_weight = grad_output.t().mm(input) if bias is not None and ctx.needs_input_grad[2]: grad_bias = grad_output.sum(0) return grad_input, grad_weight, grad_bias ``` Now, to make it easier to use these custom ops, we recommend either aliasing them or wrapping them in a function. Wrapping in a function lets us support default arguments and keyword arguments: ```Python linear = LinearFunction.apply ``` ### PyTorch NN F Activation Functions: `nn.functional.sigmoid` `relu` `softmax` Loss Functions: ### PyTorch NN Module `torch.nn.Module`: Base class for all neural network modules. Modules can contain other Modules, allowing to nest them in a tree structure. ```Python class Model(nn.Module): def __init__(self): def forward(self, x): ``` **Containers classes:** | Classes | Description | | ---- | ---- | | nn.Module | Base class for all neural network modules. Your models should also subclass this class. | | nn.Sequential(arg) | A sequential container. Modules will be added to it in the order they are passed in the constructor. | | nn.ModuleList(modules=None) | Holds sub-modules in a list. ModuleList can be indexed like a regular Python list, but modules it contains are properly registered, and will be visible by all Module methods. | Methods of nn Module:**** | Methods | Arguments | Description | | ---------------- | -------------- | -------------------------------------------------------------------------------------- | | zero_grad | () | Reset gradients of all model parameters. | | children | () | Returns an iterator over immediate children modules | | named_children | () | | | apply | (fn) | Applies fn recursively to every submodule as returned by .children() as well as itself | | parameters | (recurse=True) | Returns an iterator over module parameters | | named_parameters | () | | | buffers | (recurse=True) | Returns an iterator over module buffers | | named_buffers | () | | | to | (device) | ... | | train, eval | (mode=True) | This has effect only on certain modules, e.g. BN, dropout, etc. | *How to register parameters?* ```Python nn.Parameter(data=None) ``` A kind of Tensor that is to be considered a module parameter. *How to register buffers?* ```Python Module.register_buffer(name, tensor, persistent=True) ``` Adds a buffer to the module. This is typically used to register a buffer that should not be considered a model parameter. For example, BN's `running_mean` is not a parameter, but is part of the module's state. Buffers, by default, are persistent and will be saved alongside parameters. This behavior can be changed by setting `persistent` to `False`. The only difference between a persistent buffer and a non-persistent buffer is that the latter will not be a part of the module's `state_dict`. **State dicts:** | Methods | Arguments | Description | | --------------- | ------------------------- | --------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- | | state_dict | | Returns a dictionary containing references to the whole state of the module | | load_state_dict | (state_dict, strict=True) | Copies parameters and buffers from state_dict into this module and its descendants. If strict is True, then the keys of state_dict must exactly match the keys returned by this module's state_dict() function. | or specifics: **Linear:** Applies an affine linear transformation to the incoming data: $y=xA^T+b$. ```Python torch.nn.Linear(in_features, out_features, bias=True) ``` - input: $(*, H_\text{in})$ where $*$ means any number of dimensions including none. - output: $(*, H_\text{out})$ where all but the last dimension are the same shape as the input. - weight: the learnable weights of the module of shape `(out_features, in_features)`. The values are initialized from $U(-\sqrt{k},\sqrt{k})$ where $k=\frac{1}{\text{in features}}$. - bias: the learnable bias of the module of shape `(out_features)`. If bias is True, the value are initialized from $U(-\sqrt{k},\sqrt{k})$ where $k=\frac{1}{\text{in features}}$. **Embedding:** A simple lookup table that stores embeddings of a fixed dictionary and size. ```Python nn.Embedding(num_embeddings, embedding_dim) ``` - input $(*)$, `intTensor` or `LongTensor` of arbitrary shape containing the indices to extract - output: $(*, H)$, where $*$ is the input shape and H=embedding_dim. - weight: the learnable weights of the module of shape `(num_embeddings, embedding_dim)` initialized from `N(0,1)`. (more depths in Deep Learning) A Basic Backward Propagation: ```Python class SimpleNN(nn.Module): def __init__(self, input_size, output_size): super(SimpleNN, self).__init__() self.fc1 = nn.Linear(input_size, output_size) def forward(self, x): return nn.funtional.relu(self.fc1(x)) model = SimpleNN(10, 1) optimizer = optim.SGD(model.parameters(), lr=0.01) outputs = model(inputs) # (data loading upcoming) loss = criterion(outputs, targets) optimizer.zero_grad() loss.backward() optimizer.step() ``` ### PyTorch Utils (and other friends) **Datasets and dataloaders**: At the heart of PyTorch data loading utility is the torch.utils.data.DataLoader class. It represents a Python iterable over a dataset, with support for: - map-style and iterable-style datasets, - customizing data loading order, - automatic batching, - single- and multi-process data loading, - automatic memory pinning. ```Python DataLoader( dataset, batch_size = 1, shuffle = False, collate_fn = None, drop_last = False ) ``` Map-style datasets: A map-style dataset is one that implements the getitem and len protocols, and represents a map from indices to data samples. For example, such a dataset, when accessed with dataset$[$idx$]$, could read the idx-th image and its corresponding label from a folder on the disk. Iterable-style datasets: An iterable-style dataset implements iter and is particularly suitable for cases where random reads are improbable. For example, such a dataset, when called with iter could return a stream of data reading from a database, a remote server, or even logs generated in real time. ![[Pasted image 20231017160901.png]] Sampler: A sequential or shuffled sampler will automatically constructed based on the shuffle argument to a loader. Alternatively, users may use the sampler argument to specify a custom sampler object that at each time yields the next index to fetch. **Automatic batching:** The most common case corresponds to fetching a mini batch of data and collating them into batched samples, i.e. containing Tensors with one dimension being the batch dimension, usually the first. When batch_size is 1 and not None, the data loader yields batched samples instead of individual samples. batch_size and drop_lasts arguments are used to specify how the data loader obtains batches of dataset keys. For map-style datasets, user can alternatively specify batch_sampler, which yields a list of keys at a time. After fetching a list of samples using the indices from sampler, the function passed as the collate_fn argument is used to collate list of samples into batches. A custom collate_fn can be used to customize collation, e.g. padding sequential data to max length of a batch. In this case, loading from a map-style dataset is roughly equivalent with: ```Python for indices in batch_sampler: yield collate_fn([dataset[i] for i in indices]) ``` In particular, the default collate_fn has the following properties: - It always prepends a new dimension as the batch dimension - It automatically converts numpy arrays and python numerical values into pytorch tensors - It preserves the data structure, e.g., if each sample is a dictionary, it outputs a dictionary with the same set of keys but batched tensors as values (or lists if the values can not be converted into tensors)?! Disabled batching? Memory pinning? Multi-process loading? "A Fleged Training Loop" "How to create Dataset from folders?" ## Optimizers (need to revise them hai ek choti) Machine learning considers the problem of minimizing an objective function that has the form of a sum: $$Q(w)=\frac{1}{n}\sum_{i=1}^n Q_i(w)$$ where the parameter $w$ that minimizes $Q(w)$ is to be estimated. Each summand function $Q_i$ is typically associated with the $i-$th observation in the data set. When used to minimize the above function, a standard gradient descent would perform the following iterations: $$w=w-\eta\nabla Q(w)$$ where $\eta$ is called learning rate in machine learning. In stochastic gradient descent, the true gradient of $Q(w)$ is approximated by a gradient at a single sample: $$w=w-\eta\nabla Q_i(w)$$ As the algorithm sweeps through the training set, it performs the above update for each training sample. ![[Pasted image 20231006000027.png]] Momentum: Stochastic gradient descent with momentum remembers the update $\Delta w$ at each iteration, and determines the next update as a linear combination of the gradient and the previous update. Momentum optimization cares a great deal about what previous gradients were: at each iteration. In other words, the gradient is used for acceleration, not for speed. $$\Delta w = \alpha \Delta w - \eta \nabla Q_i(w)$$$$w = w+\Delta w$$where $\alpha$ is an exponential decay factor between 0 and 1 that determines the relative contribution of the current gradient and earlier gradient to the weight change. The momentum stems from an analogy to momentum in physics: the weight vector $w$, thought of as particle traveling through parameter space, incurs acceleration from the gradient of the loss (force). To simulate some sort of friction mechanism and prevent the momentum from growing too large, the algorithm introduces a new hyperparameter $\alpha$, simply called the momentum, which must be set between 0 (high friction) and 1 (no friction). The method was modified to use the gradient at the next point, and the resulting so-called Nesterov Accelerated Gradient was sometimes used in ML: $$\Delta w = \alpha \Delta w - \eta \nabla Q_i(w+\alpha\Delta w)$$ This works because in general the momentum vector will be pointing in the right direction towards the optimum so it will slightly more accurate to use the gradient measured a bit farther in that direction. ![[Pasted image 20231006003644.png]] If the gradient remains constant, the terminal velocity i.e., the maximum size of the weight updates is equal to that gradient multiplied by the learning rate $\eta$ multiplied by $\frac{1}{1-\alpha}$. For example, if $\beta=0.9$, then the terminal velocity is equal to 10 times the gradient times the learning rate, so Momentum optimization ends up going 10 times faster than Gradient descent. This allows Momentum optimization to escape from plateaus much faster than Gradient Descent. Gradient Descent goes down the steep slope quite fast, but then it takes a very long time to go down the valley. In contrast, Momentum optimization will roll down the valley faster and faster until it reaches the the optimum. Adaptive: Adaptive gradient algorithm is a modified stochastic gradient descent algorithm with per parameter learning rate. Informally, this increases the learning rate for sparser parameters and decreases the learning rate for ones that are less sparse. This strategy often improves convergence performance over standard stochastic gradient descent in settings where data is sparse and sparse parameters are more informative. $$s=s+\nabla Q_i(w)\circ \nabla Q_i(w)$$ $$w=w-\eta\nabla Q_i(w)\circ \frac{1}{\sqrt{s}}$$ AdaGrad often performs well for simple quadratic problems, but unfortunately it often stops too early when training neural networks. The learning rate gets scaled down so much that the algorithm ends up stopping entirely before reaching the global optimum. Adagrad slows down a bit too fast and ends up never converging to the global optimum, the RMSProp fixes it by accumulating only the gradient from the the most recent iterations as opposed to all gradients since the beginning of training. It does so by using exponential decay in the first step. $$s=\gamma s + (1-\gamma)\nabla Q_i(w)\circ \nabla Q_i(w)$$ $$w=w-\eta\nabla Q_i(w)\circ\frac{1}{\sqrt{s}}$$ where, $\gamma$ is the forgetting factor, typically set to 0.9. Adam: Adam, which stands fro adaptive moment estimation, combines the ideas of Momentum optimization and RMSProp: just like Momentum it keeps track of an exponentially decaying average of past gradients, and just like RMSProp it keeps track of an exponentially decaying average of past squared gradients. $$\Delta w=\alpha\Delta w - (1-\alpha)\nabla Q_i(w)$$$$s= \gamma w - (1-\gamma)\nabla Q_i\circ\nabla Q_i(w)$$ $$\Delta\hat{w}=\frac{\Delta w}{1-\alpha^t},~~\hat{s}=\frac{s}{1-\gamma^t}$$ $$w=w+\eta\Delta\hat{w}\circ\frac{1}{\sqrt{s+\epsilon}}$$Since $\Delta w$ and $s$ are initialized at $0$, they will be biased toward $0$ at the beginning of training, so they are boosted at the beginning of training. The momentum decay hyperparameter $\alpha$ is typically initialized to 0.9, while the scaling decay parameter $\gamma$ is initialized to 0.999. The smoothing term $\epsilon$ is usually initialized to a tiny number such as $10^{-8}$. Since Adam is an adaptive learning rate algorithm, it requires less tuning of the learning rate hyperparameter. Nadam optimization is simply Adam plus the Nesterov trick, so it will often converge slightly faster than Adam. Nadam generally outperforms Adam, but is sometimes outperformed by RMSProp. ![[Pasted image 20231006011853.png]] ## LR Scheduling A learning rate schedule changes the learning rate during learning and is most often changed between epochs/iterations. If start with a high learning rate and then reduce it once it stops making fast progress, you can reach a good solution faster than with the optimal constant learning rate. ![[Pasted image 20231006171749.png]] Power scheduling: Set the learning rate to a function of the iteration number $t$: $\eta(t) = \frac{\eta_0}{(1+t/k)^c}$. The initial learning rate $\eta_0$, the power $c$ typically set to 1 and the steps $s$ are hyperparameters. The learning rate drops at each step, and after $s$ steps it is down to $\eta_0/2$. After $s$ more steps, it is down to $\eta_0/3$. Then down to $\eta_0/4$, then $\eta_0/5$, and so on. Exponential scheduling: Set the learning rate to $\eta(t)=n_0 0.1^\frac{t}{s}$. The learning rate will gradually drop by a factor of 10 every $s$ steps. Cosine scheduling: ## Weight decay ## FP16 training ## PyTorch Geometric torch_geometric.data: Data: A Data object describes a homogeneous graph. The data object can hold node-level, link-level and graph-level attributes. In general, Data tries to mimic the behavior of a regular Python dictionary. In addition, it provides functionality for analyzing graph structures, and provides basic PyTorch tensor functionalities. ```python Data(x, edge_index, edge_attr, y, pos); (each optional) Parameters: x: node feature matrix with shape (num_nodes, num_node_features) edge_index: graph connectivity in COO formatt with shape (2, num_edges) edge_atrr: edge feature matrix with shape (num_edges, num_edge_features) y: graph level or node level ground truths with arbitrary shape pos: node position matrix with shape (num_nodes, num_dimensions) ``` Batch: A data object describing a batch of graphs as one big disconnected graph ```python pyg.data.Batch().from_data_list([graph1, graph2]) to_data_list() ``` Dataset: Dataset base class for creating graph datasets, probably is a collection of Data i guess, with ofc extra information ```python Dataset(root, transform, pre_transform) transform vs pre_transform? as we access from disk vs before saving to disk Properties: len(): no of data objects [idx]: access the data obj at index idx data: gives the collection of datasets as one large data object num_classes: the number of classes in the dataset (probably says what y encodes) Example: name = 'Cora' transform = pyg.transforms.Compose([ transforms.RandomNodeSplit('train_rest', num_val=500, num_test=500), transforms.TargetIndegree(), ]) cora = pyg.datasets.Planetoid ( './data', name, pre_transform=transforms.NormalizeFeatures(), transform=transform ) ``` nn torch ko jastai ta hola ni: Linear Convolutional: ``` ``` Transforms: Random node split performs a node-level split random split by adding train_mask, val_mask and test_mask attributes to Data: ```python RandomNodeSplit('train_rest', num_val=500, num_test=1000) (after this the data will have extra property with these names) ``` TargetIndegree saves the globally noramlized degree of target nodes: node degree/all others degree: $$u(i,j)=\frac{\text{deg}(j)}{\text{max}_{v\in V}\text{deg}(v)}$$ (many graphs wont have edge attributes so this puts something into them) ``` TargetInDegree() ``` The normalizefeatures does 'row normalizes' rey not sure what this is though?? Loader: A data loader which merges data objects from a torch_geometric.data.Dataset to a minibatch. ```python DataLoader(dataset, batch_size=1, shuffle=True) ``` The each element of dataloader is now a databatch object. ## Fast AI >Directly run the following fastai code to see what it does then only comes the theory part: from fastai.vision.all import * path = untar_data(URLs.PETS)/'images' dls = ImageDataLoaders.from_name_func( path, get_image_files(path), valid_pct=0.2, seed=42, label_func=lambda x: x[0].isupper(), item_tfms=Resize(224) ) learn = vision_learner(dls, resnet34, metrics=error_rate) learn.fine_tune(1) 1. A dataset called oxford iiit pet dataset contains 7349 images of cats and dogs from 37 breeds is donwloaded to gpu server and extracted 2. A pretrained model that has already been trained on 1.3 million images using a compeition winning model will be donlwoad from the internet 3. The pretrained model will be finedtuned using the latest advances in transfer learning to create a model that is specially customized for recognizing cats and dogs Note: if you run 1 and 2 again then it will use the donwloaded dataset and model but train again on the GPU What goes under the hood? -First is import all the functions and classes need to create variety of computer vision models -The untar downloads a standard dataset from the fast.ai datasets collection to your server, extracts it if not previously and returns a Path object with the extracted location -Third line tells fastai what kind of dataset we have and how it is structured here is ImageataLoaders >path is where our data is >get_image_files gets images recursively from the path >valid_pct is for validation ratio >seed >label_func is parent folder name as cat is uppercase >Resize to 224 pixel is a transfrom that is applied automatically to exs Note: The metrics is calculated using validation set and never the training set -Forth is create a vision architecture that take in structured datasets and is pretrained ImageNet resnet with error rate quality measurement on validation set which is not same as loss function -Fifth tells fastai how to finetune (not fit) the model by passing number of epochs to run >Use one epoch to fit just those parts of the model necessary to get new random head to work correctly with your dataset >Use number of epochs requested when calling the method to fit the entire model, updating the weights of the later layers (especially the head) faster tahn the earlier layers s >For one line definition: instead of telling the computer the exact steps required to solve a problem, show it examples of the problem to solve, and let it figure out how to solve it itself from its experience Feedback loops: >A critical insight comes from considering how a model interacts with its environment which can create feedback loops: 1. A predictive policing model is created based on where arrests have been made in the past, which is not predicting actual crimes so there is partial biases 2. Officers then use that to decide where to focus their policing activity, resulting in increased arrests in those areas 3. Data on these additional arrests would then be fed back to retain future versions of the model Note: This is a positive feedback look the more the model is used, the more biased the data becomes making the model even more biased and so forth :Overfitting is single most important and challenging issuse as it is easy to create a model that does a great job at making predictions on the exact data it has been tarined on but much harder to make accurate predictions on data the model has never seen before: When does overfitting occurs and how to avoid it? -If you train for too long with not enough data you see the accuracy of your model on validation set start ot get worse -Use avoidance only after you have confirmed that overfitting is occuring ie if you have observed the validation accuracy getting worse during training Blackbox neural nets: -Research exists showing how to deeply inspect deep learning models and get rich insights from them Is this for image only? -A lot of things can be represented as images which means an recognized can learn to complete may tasks for instance, a sound can be converted to a spectrogram which is a chart that shows amount of frequency at each time in an audio file -A time series can easily be converted into an image by simply plotting the time series on a graph -However it is often a good idea to try to represent your data in a way that makes it as easy as possible to pull out the most important components; in a time series, things like seasonality and anomalies are most likely to be of interest -Other instances are mouse movements and binary files converted into images Note: If the human eye can recognize categories from the image, then a deep learning model should be able to do so too Whats more than classifying images? -Another important is localizing object in a picture for autonomous vehicles -Following is fastai code using a subset of the CamVid dataset of whose flow is same as before: path = untar_data(URLs.CAMVID_TINY) dls = SegmentationDataLoaders.from_label_func( path, bs=8, fnames = get_image_files(path/"images"), label_func = lambda o: path/'labels'/f'{o.stem}_P{o.suffix}', codes = np.loadtxt(path/'codes.txt', dtype=str) ) learn = unet_learner(dls, resnet34) learn.fine_tune(8) -See the results of the model using: learn.show_results(max_n=6, figsize=(7,8)) Okay enough about images? any text? -Can generate text, translate automatically from one language to another, analyze comments and much more -To train a model that can classify the sentiment of a movie review better than anything that existed in the world five years ago: -Dataset is IMDb large movie review: dls = TextDataLoaders.from_folder(untar_data(URLs.IMDB), valid='test') s learn = text_classifier_learner( dls, AWD_LSTM, drop_mult=0.5, metrics=accuracy ) learn.fine_tune(4, 1e-2) -Predict the reviews as: learn.predict("I really liked that movie brother") I want tabular man? -Usually no pretrained available for this task so use fit instead of fine tuning: -The process is similar as the following code predict whether a person is a high-income earner based on their socio-economic background: -Dataset is using Adult from a paper that scales up the accurarcy of Naive-Bayes Classifiers from fastai.tabular.all import * path = untar_data(URLs.ADULT_SAMPLE) dls = Tab Note: Had to tell which columns are categorical ie contain values that are one of a discrete set of choices such as occupation versus continuous ie contain a number that represents a quantity such as age One more: -Recommendation systems are important particularly in ecommerce so following code predicts movies people might like based on their previous viewing habits using the MovieLens dataset: from fastai.collab import * path = untar_data(URLs.ML_SAMPLE) dls = CollabDataLoaders.from_csv(path/'ratings.csv') learn = collab_learner(dls, y_range(0.5, 5.5)) learn.fine_tune(5) learn.show_results() -Note using a pretrained model for the same reason we didnt for the tabular model but uses fine_tune anyway; its best to experiment fine_tune vs fit_one_cycle to see which works best for your dataset VALIDATION SETS: -If we trained a model with all our data and then evaluated the model using the same data we would not be able to tell how well our model can perform on the data it hasnt seen -Susequent versions of the model are indirectly shaped by us having seen the validation data; just as the automatic training process is in danger of overfitting the training data we are in danger of overfitting the validaiton data through human trial and error and exploitation -Solution is to introduce another level of more highy refined data: test setwhich can only be used to evaluate the model at the very end of our efforts Note: Test and validation tests should have enough data to ensure that you get a goog destimate of your accuracy, if you are creating a cat decetor need 30 cats in validaiton sets, if you hvae dataset with thousands of items using 20% of the set may be more than you need "A KEY PROPERTY OF THE VALIDATION AND TEST SETS IS THAT THEY MUST BE REPRESENTTATIVE OF THE NEW DATA YOU WILL SEE IN THE FUTURE" 1. Want more than just randomly grap a fraction of your original dataset 2. If you are looking at time series data choosing a random set will be easy and no representative of most business use cases, so you will want to choose a continouous section for instance, two weeks or last month of avaialble data 3. Another common is when you can easily anticipate ways the data you will be making predictions for in production may be qualitatively differnet from the data you have to train your model with; ex distracted driver compeition where images are of same person in different positions Production tips: -Keep an open mind to the possiblity that deep learning might solve part of your problem with less data or complexity than you expect -Goal is not to find the perfect dataset but to get started and interate from there -Best to first start by finding example online of some thing that somebody has had good results with and that is at least somewhat similar to what you are trying to achieve by converting your data into a similar format to what someone else has used before such as creating an image from your data State-of-arts: 1. Computer vision -Major works around recogniztion, detection and segmentation -Generally not good at recognizing images that are significantly different in structure from those used to train the model for ex: may be only black and white or only hand-written images in train set -No general way to check which types of images are missing in your training set -Labelling can be slow and expensive, one particularly helpful is to synthetically generate variation of input images by rotating or chaning their brightness ie. data augmentation 2. Text: -Good at classifying both short and long documents based on categories, sentiment (pos or neg), author, source website, and so forth -Also good ate generating context-appropriate text such as replies to social media posts, and imitatint a particular authors style -However is not good at generating correct responses so dont have a reliable way to combine a knowledge base of medical info with a deep model for generating medically correct natural language responses -Many applications such as translation from one lang to another, summarize long documents, find mention of a concept of interest and more -Well could include completely incorrect information but the performance is good enough to be used in current systems 3. Combining text and images: -Ability to combine text and images into a single model is generally far better than most people intutively expect -Can train on input iamges with output captions to generate suprisingly appropriate captions automatically for new images: with no gurantee that these captions will be correct -So recommended to be used as part of a process in which the model and a human user interact closely -Text to images? 4. Recommendation systems: -Are just really a special type of tabular data that generally have a high cardinaltiy categorical variable representing users, and another one representing products or something similar -As deep models are good at handling high cardinality categorical variables, they are quite good at handling recommendation systems 5. Other data types: -Ofther domain specific data types fit nicely into existing categories, for instance, protein chains look a lot like natural langugage documents, in that that they are long sequences of discrete tokens with complex relationships and meaning throughout the sequence -Sounds can be represented as spectrograms which can be treated as images; turn out really well on spectrograms Approach to deep learning by Jeremy: 1. Start with considering your objective 2. Think about what actions you can take to meet that objective 3. What data you have or can acquire that can help 4. Build a model that you can use to determine the best actions to take to get the best results in terms of your objective Gathering data: -For many types of projects you may be able to find all the data you need online through services from fastcore.all import * from fastai.vision.all import * from duckduckgo_search import DDGS from fastdownload import download_url #Get urls for your search: urls = L(DDGS().images(' ... ')).itemgot('image')[:100] #Check one of the url: dest = '....jpg' download_url(urls[0], dest, show_progress=False) im = Image.open(dest) im.to_thumb(256 ,256) #Download all the images: path = Path('images') cat = '...' dest = (path/cat) dest.mkdir(exist_ok=True, parents=True) download_images(dest, urls=urls) #Delete the failed images: failed = verify_images(get_image_files(path)) failed.map(Path.unlink) #Make something called datablock structure that contains train and valid sets data = DataBlock( #inputs are images and outputs are categories: blocks=(ImageBlock, CategoryBlock), #use get_image_files which returns a list of all image files in a path get_items=get_image_files, #train and test set splitter=RandomSplitter(valid_pct=0.2, seed=42), #labels are just folder names get_y=parent_label, #resize the images item_tfms=[Resize(192)] ) #Make dataloader out of the datablock dls = data.dataloaders(path) dls.show_batch(max_n=6) #Pretraining ImageNet resnet34 model: learn = vision_learner(dls, resnet34, metrics=error_rate) learn.fine_tune(3) #Prediction using the model: learn.predict(PILImage.create('....jpg')) Data vs dataloaders: -Dataloaders is a thin class that just stores whatever DataLoader objects you pass to it and makes them available as train and valid for the models -To turn our donwloaded data into a DataLoaders object need to tell fastai at least four things 1. What kinds of data we are working with (blocks) 2. How to get the list of items (get_items) 3. How to label these items (get_y) 4. How to create the validation set (splitter) -Upto now pretraining was eased with data structure that happen to fit into those pretrained model, for when not available, fastai has an extermely felxible system called the datablock API with which can customize every stage of the creation of your DataLoaders -Need to add a transform that will resize these images to the same size, so images can fit into a tensor (item_tfms) -By default Resize crops the images to fit a square shape of the size requested using the full width or height; or alternatively could squish or stretch them data = data.new(item_tfms=Resize(192, method='squish')) -All these seem wasteful instead what we normally do in practice is to randomly select part of the image and them crop to just that part data = data.new(item_tfms=RandomResizedCrop(192, min_scale=0.3)) -The random crop is a specific example of a more general technique called data augmentation which refers to creating random variations of our imput data, such that they appear different but do not change menaing of the data -For natural images: data = data.new(item_tfms=Resize(192), batch_tfms=aug_transforms(mult=2)) Using the model to clean the data: -To visualize the mistakes the model is making can create a confusion matrix which is calculated using the validation set interp = ClassificationInterpretation.from_learner(learn) interp.plot_confusion_matrix() -Helpful to see where exactly are ours error occuring to see whether they are due to a dataset problem or a model problem, to do which can sort our images by their loss, which is high when the model is incorrect especially but also confident of its incorrect answer or if its correct but not confident of its correct answer (must do this) interp.plot_top_losses(6) -The general intuitive approach to data cleaning is to do it before you train a model but a model can help you find issues more quickly and easily so we normally prefer to train a quick and simple model first, and then use it to help us with data cleaning -fastai includes a handy GUI for data cleaning that allows you to choose a category and the training versus validation set and view the highest loss images in order along with menus to allow images to be selected for removal from fastai.vision.widgets import ImageClassifierCleaner cleaner = ImageClassifierCleaner(learn) cleaner Note: After cleansing need to recreate datablock to retrain the model for idx in cleaner.delete(): cleaner.fns[idx].unlink() Turning your model into an online application: -Once model is ready, save it to copy to the production server to save which is the export method: learn.export() -To create our inference learner from the exported file, we use load_learner as: path = Path() learn = load_learner(path/'export.pkl') -Also can access DataLoaders object as an attribute of the learner: learn.dls.vocab The notebook app: -Can create a complete working web app using nothing but Jupyter notebooks using ipywidgets and voila -IPython widgets are GUI components that bring together JS and python functionality in a web browser and can be created and used within a Jupyter notebook -Voila exists for making applications consisting of IPython widgets available to end users without them having to use Jupyter at all 1. Create a button to upload images for classification: from fastai.vision.widgets import widgets btn_upload = widgets.FileUpload() btn_upload 2. Create a button to run the inference on the models: btn_run = widgets.Button(description='Classify') btn_run 3. Create a cell for images under classifiton process: out_pl = widgets.Output() out_pl 4. Create a cell for displaying the propbabilities: lbl_pred = widgets.Label() lbl_pred 5. Create a function that runs when click on the classify: def on_click_classify(change): img = PILImage.create(btn_upload.data[-1]) out_pl.clear_output() with out_pl: display(img.to_thumb(128,128)) pred,pred_idx,probs = learn.predict(img) lbl_pred.value = f'Prediction: {pred}; Probability: {probs[pred_idx]:.04f}' btn_run.on_click(on_click_classify) -First create an img out of the last uploaded image -Clear the old output to make room for display -Make prediction and display the probabilities 6. Can put all the widgets in a vertical box to complete GUI: VBox([ widgets.Label('Select your bear!'), btn_upload, btn_un, out_pl, lbl_pred ]) ## SKLearn CODING: from sklearn.decomposition import PCA pca = PCA(n_components = 2) train_set_2d = pca.fit_transform(train_set) pca.explaind_variance_ratio_ #gives the percentage of data points in each axis >To choose the right number of dimensions for just decomposition do: pca = PCA pca.fit(train_set) cumsum = np.cumsum(pca.explained_variance_ratio_) d = np.argmax(cumsum >= 0.95) + 1 >Now we could do n_components = d which preserves 95% of the variance CODING: from sklearn.tree import DecisionTreeClassifier tree_clf = DecisionTreeClassifier( criterion='gini', max_depth=3, min_samples_split=3 #how many samples must be in a node to consider split ) tree_clf.fit(train_set, train_set_labels) predictions = tree_clf.predict(train_set) probabs = tree_clf.predict_proba(train_set) tree_clf.feature_importances_ #provides how each feature contribute to making decision CODING: from sklearn.naive_bayes import BernoulliNB naive_clf = BernoulliNB() naive_clf.fit(train_set, train_labels) naive_clf.predict(train_set) naive_clf.predict_proba(trainset) CODING: from sklearn.neighbors import KNeighborsClassifier neighbor_clf = KNeighborsClassifiers( n_neighbors = 5, p = 2, metric = 'minkowski' ) [there is also KNeighborsRegressor] K-means clustering: -To identify groups of similar looking objects, just like in classificaiton each instance gets assigned to a group with unsupervision -Kmeans designate k blobs and assign data points to either one of them, first randomly initializes the centroids of each blob and points are assigned by the distance metric of which centroid they are close to -Can be done iteratively by assiging points to the closest centroid value blob, then calculating centroid of points into the blob and so on, centroid is just mean of all the points -Does not necessarily go to global minimum, may be stuck on local minimum as well sometimes well -If there are not absolute blobs then quality of this shit decreases significantly CODING: from sklearn.cluster import KMeans kmeans = KMeans(n_clusters=5, init=[[centroid 1], [centroid 2], [], []], n_init=1) kmeans.fit(train_set) predictions = kmeans.predict(train_set) #returns an array of numbers belonging class kmeans.cluster_centers_ #is the list of centroid found kmeans.transform(train_set) #for each example provides distance from each centroid so essentially decomposes into k-dimension Regression: -Linear regiresson where each example x is an n-element vector, out hypothesis space is the set of functions of the form h(x vector) = w0 + w1x1 + ... + wnxn -Let N be number of training examples and n be features (w0...wn) -In linear algebra power and introduce x0 = 1 h(x vector) = (w) dot product (x) -We find w by minimizing what we call the expected values of loss over examples -Gauss showed that if yj values are normally distributed noise, then the most likely values of w's are obtained by using L2 loss, minimzing the sum of squares of the errors, if the values have noise that follows a Laplace distribution then L1 loss is appropriate -If we have some w vector then its loss depends on examples X vectors and y labels L(w vector) = sum over i (yi'- yi)^2/2N (expected loss) [The divide by 2 is for beaut] -To minimize we need the graidnet ofc: dL(w vector)/dwi = (yi' - yi)xi -Vectorially also has a nice form: grad L = X^T (Xw) [X matrix made by arranging xvectors rowwise] [Tempting to expand the inverse right? but the X is not necessarily invertible bro] -Then later: -Hence training takes O(n^3) because there are matrix multiplication invovled -Another way is by SVD as follows: X = s@v@d then, X+ = d.T@v+@s.T -To calculate v+, take v and set zero to all values smaller than a tiny threshold value, then replaces all the nonzero values with their inverse and finally tranpose the resulting matrix -So the inversion step in original has been replaced by finding v+ which takes O(n^2) as compared to O(n^3) where n is the number of features so the scaling with features is obvious here CODING: from sklearn.linear_model import LinearRegression lin_reg = LinearRegression( copy_X: True, //to overwrite or not fit_intercept: True //whether to set bias to zero positive: False //are all attributes weights positive? n_jobs: None // How many CPUs to use ) lin_reg.fit(train_set, train_set_labels) predictions = lin_reg.transform(train_set) Fast fourier transform: But before that: -Linear models requires dataset to be: >Fully numericals >No missing data at all >Better to minimize the outliers >Adding and removing attributes on coorelation 1. OneHotEncoder() to convert non numerics into numerics 2. SimpleImputer(strategy='median') for placing out the missing values 3. Custom 4. Custom >They are put through the ColumnTransformer to generate numpy arrays of trainset Gradient descent: -Visuailze weights vs cost graph which is always convex if you use L2 so another reason to use L2, the gradient point to steepest ascent so we need negative of it w(new) = w(old) + eta * direction of steepest descent >Direciton of steepest descent depends on current loads of examples and their output with old weightss -The size and direction of next step is given by: w(new) = w(old) - eta * graident of MSEs -The use of whole examples at once is called the batch graident descent, a faster variant is called stochastic gradient descent; -It randomly picks up a random instance in the training set at every step and computes the graidnets based only on that single instance -Much less regular than batch insstead of gently decreasing will bounce up and down decreasing only on average, overtime it ends up very close to the minimum but once it gets there it will continue to bounce around, never settling down, so once it stops, the final parameter values are good, but not optimal -Now there are few hyperparameters to vary >eta is the learning rate >number of iterations to decide >tolerance level of the last epoch graident >learning schedule by which the eta varies over epoches -Each provided and implemented in SGDRegressor of sklearn -A gradient descented depends on distance so scaling is better you know CODING: from sklearn.linear_model import SGDRegressor sgd_reg = SGDRegressor( eta0 = 0.01, max_iters = 1000, tot = 0.001 learning_rate = 'invscaling' ) sgd_reg.fit(train_set, train_set_labels) predictions = sgd_reg.transform(train_set) >The gradient descent stops on the following condtions: 1. If reaches max_iter 2. If reaches tol level of tolerance fo n_iter number of iterations which could use validation set if you do early_stopping=True Polynomializing: -Linear ta alli nahuna sakxa haina halka quadratic or cubical huna sakxa testo case ma attributes lai convert garera higher degrees ko ni add garni paryo CODING: from sklearn.preprocessing import PolynomialFeatures poly_features = PolynomialFeatures(degree=2, include_bias=False) X_poly = poly_features.fit_transform(X) which adds features upto degree in addition to the existing ones Learning curves: >Can do cross vliadation to get an estimte of a models general performance, if a model performs well on the trainign data but generlized poorly on cross validation then is overfitting, if poor on both then underfitting >Another way is to look at the learning curves: plots of the models performance on the trainiigns set and validatio set as a function of the traning set size, to generate the plots simply train the model several times on different sized subsets of the training data CODING: from sklearn.model_selection import learnign_curve train_sizes, train_errors, validation_errors = learning_curve( estimator = <trained model name here> X = train_set, y = train_labels, cv = 3, train_sizes = [1000, 2000, 3000, 40000, 5000] scoring = 'neg_mean_squared_error' ) train_errors = [np.average(np.sqrt(-x)) for x in train_errors] validation_errors = [np.average(np.sqrt(-x)) for x in validation_errors] scores = pd.DataFrame( { "train_sizes": train_sizes, "train_errors": train_errors, "validation_errors": validation_errors } ) -When both curves have reached a plateau and they are close and fairly high means they are underfitting; they cannot improve if new data is added so required complex models -If there is gap between curves means performs significantly better on trianing than validation, if new data is added would slowly improve Regulaizarion: -To constraints the weights of the model add a direct punishment to higher weights in the model itself -Ridge adds squared, lasso absolute, which are scaled by a factor of alpha, increasing which increases the degree of punishment -Which to use depends on specific problems, but L1 temds to produce a sparse model that is often sets many weights to zero, effectively declarin the corresponding attributes to be completely irrevelant -This is because for some value of lamda the compleixt(h) <= c where c realtes to lamba, now for L1 regulartion the curve of complexity <= c is a square whose vertiex are on axes equatoin is probalby |x|+|y|=1 swile for L2 it is a circle, now the weight must lie on to this box or circle or whatever in higher dimeions CODING: from sklearn.linear_model import Ridge, Lasso Lasso(alpha=1) Ridge(alpha=1) OR in gradient descent: SGDRegressor(..., alpha=1, penalty='l2') FLOW: -Requires fully numerical and complete dataset and probably no outliers -To add polynomializing features or not brother -always do some ridging -To do analytical or gradient descent >To choose alpha we could can do grid search on the hyperparameters that try all the combinations and find you the best one: CODING: ---------------------------- Linear classifiers: -Can be made for classification using the straight line as a decision boundary that separates the two classes, data need be linearly separable h(x vector) = sigma (x dot w) -The sigma function makes sense to be a heaviside but train garna mildena so we use something called sigmoid whose output can be interpreted as probabliy that x vector falls in class 1 -Loss function is given as: L(w vector) = expected cross entropy between our predicted and actual class [earlier was expected L2 loss over difference between predited and actual value] L(w vector) = sum over all samples later divided by N [distribution of being in class 0 log (1/same predcited) + distribution of being in class 1 (1/pred)] As compact, = sum over i: -yi*log(sigma(yi'))-(1-yi)(1-sigma(yi'))]/N [which when solving for graident descent taking derivative has no closed form for finding w*] [from future: i think the closed form is same as that of L2 loss expect for that 2] -The gradeint update rule is based on neurons that wire together fire together form CODING: from sklearn.linear_model import SGDClassifier sgd_clf = SGDClassifier( loss = 'log_loss' eta0 = 0.01, max_iters = 1000, tot = 0.001, learning_rate = 'invscaling' alpha=1, penalty='l2' ) sgd_clf.fit(train_set, train_set_labels) sgd_clf.predict_proba(train_set) sgd_clf.predict(train_set) where decision boundary is 0.5 Softmax classifiers: -Can be generalized to support multiple classes directly without having to train and combine mulitple binary classifiers -Now there are many w vectors which will tell the probabilty of x vector falling into that class -Hence, h_i(x vector) = exp(x dot w) / (sum over j exp(wj dot x)) This is h function of (x, and every w there is) is the probabilty that x vector falls into ith class where tyo exp wala lafda xa ni tyo chai generalized sigmoid jastai ho bhanda hunxa -Now highest probablity bhako lai yei class ko ho bhanda huni bhayo -Loss function: L(w vectorS) = expected cross entropy using distribuiton for predcited and actual class: = sum over examples: [sum over k: distrution of being in class k * log(1/predicted class distriution for k)] L(w vectorS) = sum[-sum on each class k: actual_y*log(sigma(Xw_i)]/N -For two classes is equivalent to logistic regression so is a pretty good generalization -Notice that the loss function is actually expected cross entropy where actual distribution is 0s and 1s from our dataset and predicted is our function predcition, we try to minimize it as much as possible so we are certain about the inforamtion CODING: -There is another logistic class which uses other weird form of learnings which probably i wont understand from sklearn.linear_model import LogisticRegression softmax_reg = LogisticRegression(multi_class='multinomial') softmax_reg.fit(train_set, train_set_labels) softmax_reg.predict_proba(train_set) In summary, the linears fits and their polynomial extensions: 1. Hot encode the categoricals 2. Impute missing values 3. Minimize the outliers (which could cause overfitting) 4. Add or remove attributes based on correlations 5. Polynomialising the attributes if you see the pattern 6. Standard scale the attributes 7. SV or Linears and their regulaizarions with SGD or not [Softmax avaialble for LCs, kernel trick for SVCs] 8. Cross validate and plot the learning graph or the precision-accuracy tradeoff 9. Grid search if there are any paramters need to tune in CODING: #sequential API os keras models = keras.models.Sequential() #just calculates reshape(-1,1): models.add(keras.layers.Flatten(input_shape=[28,28])) #hidden dense layers with respective neuron capacitites: model.add(keras.layers.Dense(300, activation="relu") model.add(keras.layers.Dense(100, activation="relu") //Here each layer handles a matrix of weights (one vector of weights for each neuron) and also a vector of bias term (one per neuron) and calulates following: h(W,b) (X vector from previous layer) = phi (XW + b) //bias written separately as is for the deep learning //Except ofc the input layer which takes (non vector entity) and thrwos out the vector one Also by directly passing: model = keras.models.Sequential([ keras.layers.Flatten(input_shape=[28, 28]), keras.layers.Dense(300, activation="relu"), keras.layers.Dense(100, activation="relu"), keras.layers.Dense(10, activation="softmax") ]) >Check model review by: model.summary() which gives following info: 1. Paramters of each layer clcaulated by number of inputs * number of neurons + number of neurons which is very large for 300 neurons which poses a risk of overfitting >To refer to each of the layers do: model.layers[index] >Can do set_weights() and get_weights() to each layer: weights, biases = layer_name.get_weights() return an numpy array of same NOTE: The dense layer initializes the weights randomly but biases are put to zero >Can do our own initilizement though later >The certain number of paramters we saw is due to the fact that we did input_shape but if you dont, Keras will simply wait until it knows the input shpae before it actually builds the modoel, this happens when you actually feed it actual data or when you call its build() >Until the model is really built, layers wil not have any weights, and you will not be able to do certian things (such as print the model summary or save the model) -Model is created meaning that we have specified the input features and acitvations but we have not given its loss function and trianing method -To do which is to COMPILE the model: model.compile(loss="sparse_categorical_crossentropy", optimizer="sgd", metrics=["accuracy"]) Here note: >We are using sparse cause the labels are one hot encoded: our output is a 10 elements of softmax while the labels are like [1] for coat and [2] for jacket and so onsss >Sigmoid use garera multilabel classificaiton garni bhane chai we would be using the binary crossentorpy loss function >Finally building and compiling model now time to train: history = model.fit(X_train, y_train, epochs=20, validation_data=(X_valid,y_valid)) //instead of actuallly passing a validation set you could use validation_split=0.1 that kears will use >What if like the previosu problem? the dataset was skewed? some arguments are there to pass >Return the history object containing -The training paramters (history.params) -The list of epochs it went through history.epochs -The dictionary containg the loss and metrics it measured at the end of each epoch on the trianing set and on the validation set >Can convert into a DF and plot to see the loss and accuracy as a fucntion of epoches >Then you find from the curve that you could do better by increasing the epoch, in which case you could just call fit() since Keras continues trianing where it left off ........ >Now evaluate on test sets: >model.evaluate(X_test, y_test) to check on the data set >Get your predictions brother: y_pred = model.predict(X_new) returns an array with 1 in the place of its actual calss >Regression: same way Functional API: -To connect all or part of the inputs direeclty to the output layer which makes it possible for the neural network to learn both deep patterns (using the deep path) and simple rules (through the short path) -In contrast, a regular MLP forces all the data to flow through the full stack of layers thus simple patterns in the data may end up being distorated by this sequence of transformations -For housing problem: input = keras.layers.Input(shape=X_train.shape[1:]) //need to create as we may have multiple inputs, I see is that we may be sending scalar or vector arrays so better to get them in same format hidden1 = keras.layers.Dense(30, activation="relu")(input) //Tell functional API how to connect the layers together, no actual data is being processed, less neurons than classifiers cause data is noisy they say and we dont want to overfit hidden2 = keras.layers.Dense(30, activation="relu")(hidden1) //pass the hidden layer to second hidden layer concat = keras.layers.Concatenate()[input, hidden2] //inputs are coming into some hidden layers and also directly and need them to concatenate for the reason mentioned above output = keras.layers.Dense(1)(concat) //the concat layer will finally go into the output one model = keras.models.Model(inputs=[input], outputs=[output]) //We had used models.Sequential in sequential api here we using Model by passing input layer and output layers #Now everything is same as earlier compiling and fitting and predicting: MULTIPLE INPTUS: -Here we had all the features input going into the output through the same layers but ometimes you want some features to go through deep layers and others direclty to the output: -They could be overlapped as well -Completely different example where we want to send 5 features (0 to 4) through deep path and 6 features through the wide path (features 2 to 7): input_A = keras.layers.Input(shape=[5,]) input_B = keras.layers.Input(shape=[6,]) hidden1 = keras.layers.Dense(30, activation="relu")(input_B) hidden2 = keras.layers.Dense(30, activation="relu")(hidden1) concat = keras.layers.concatenate([input_A, hidden2]) //inputA features is directly coming while inputB features are coming through a deep layer output = keras.layers.Dense(1)(concat) model = keras.models.Model(inputs=[input_A, input_B], outputs=[output]) MULTIPLE OUTPUTS: -Also cases when you want multiple outputs, like you want to locate and classify main object in a picture which is both regression for coordiantes and classification task -Similarly may have multiple independnet tasks to perform based on same data, could be done by training neural net for each purpose but in many cases get better result as neural net can learn faetures in the data that are useful across tasks -Another is regularization technique for eg you may want to add some auxilary outputs in a neural net architecture to ensure that the underlying part of the network learns something useful on its own without underlying on rest of the network, means same thing predicting but from different route # same as above multi inputs output = keras.layers.Dense(1)(concat) aux_output = keras.layers.Dense(1)(hidden2) model = keras.models.Model(inputs=[input_A, input_B], outputs=[output, aux_output]) -So what to do with compile one? -We need to have a complete global loss function which we choose to be weighted version of losses from individual outputs as: model.compile ( loss = ["sparse...", "sparse..."], loss_weights = [0.9, 0.1] //Give 0.1 to aux output as it is for regularization optimizer = "sgd" metrics=["accuracy"] //which comes out for each output layers ) >Now fitting: model.fit( [train_X_A, train_X_B], [train_y, train_y], epochs=20, validation_data ([valid_X_A, valid_X_B], [valid_y, valid_y]) ) >Same way of evaluation: Subclass APIing: -Some models involve loops, varying hsapes, conditional branching, and other dynamic behaviours, for such is subclsses -HAHAHAHA Saving a model: -Saving and loading a trained model: model.save("my_keras_model.h5") model = keras.models.load_model("my_keras_model.h5") Callbacks: -Go into with the fit call and are called at the end of each epoch 1. As a early stopping which interrupts training when measures no progress on the validation set for a number of epochs (defined by the patience argument), and it will roll back to the best model, if not enabled uses the recent model early_stopping_cb = keras.callbacks.EarlyStopping ( patience=10, restore_best_weights=True ) history = model.fit(..., callbacks=[early_stopping_cb]) whichs waits for 10 epochs for which until there is no improvement and also restore the best one 2. If we use validation set during trianing, and want to write the model if it is best so far then, checkpoint_cb = keras.callbacks.ModelCheckpoint("my_keras.model.h5") history = model.fit(..., callbacks=[checkpoint_cb]) VISUALIZATION: - Fine-tuning hyperparamters: -Simplest way of sklearn gridsearchcv or randomizedsearchcv: -First step is to wrap the Keras models in objects that mimic regular scikit-learn regressors which we start by creating a function that will build and compile a Keras model given a set of hyperparamters: #Here looks like each layer will have same number of neurons: def build_model(n_hidden, n_neurons, learning_rate, input_shape): model = keras.models.Sequential() model.add(keras.layers.Flatten(input_shape=input_shape)) for layer in range(n_hidden): model.add(keras.layers.Dense(n_neurons, activation="relu")) model.add(keras.layers.Dense(10, activation="softmax")) model.compile(loss="sparse_categorical_crossentropy", optimizer="sgd") return model keras_clf = keras.wrappers.scikit_learn.KerasClassifier(build_model) **Linear algebra**: ```Python np.diag(ndarray) np.outer(ndarray, ndarray) np.inner(ndarray, ndarray) ndarray@ndarray combines consecutive vector space transformations np.linalg.matrix_power(ndarray,n) np.linalg.inv(m) reverses a square transformation if no information was lost np.linalg.matrix_rank(m) is the dimension on the resulting vector space spanned by the transformation np.linalg.det(m) is the scaling factor of hypercube for square transformations q,u = np.linalg.qr(m) qr decomposes square X into QU where Q is a orthogonal such that Q.T is the inverse of Q [m = q@u] q,a = np.linalg.eig(m) np.linalg.eighvals(m) eig decomposes square X into QAQ'' where Q is a transformation whose columns are eigvecs: m = q@diag(a)@inv(q) q,a = np.linalg.eigh(m) np.linalg.eigvalsh(m) special case for the symmetric or hermitian matrix where Q is the orthogonal: m = q@diag(a)@q.T s,v,d = np.linalg.svd(m) svd decompsoes X into vaw.T where v and w are eigen decomposition of L-R symmetric matrices: m = s@diag(v)@d ```